Note
Click here to download the full example code
Custom score matrices#
This tutorial shows how to train a custom score matrix for NBLAST.
The core of the NBLAST algorithm is a function which converts point matches (defined by a distance and the dot product of the tangent vectors) into a score expressing how likely they are to have come from the same cell type. This function is typically a 2D lookup table, referred to as the "score matrix", generated by "training" it on sets of neurons known to be either matching or non-matching.
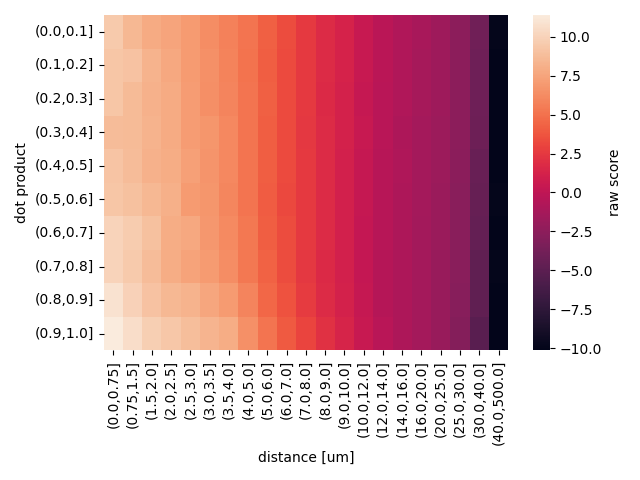
NAVis provides (and uses by default) the score matrix used in the original publication (Costa et al., 2016) which is based on FlyCircuit, a light-level database of Drosophila neurons. Let's quickly visualize this:
import navis
import seaborn as sns
import matplotlib.pyplot as plt
smat = navis.nbl.smat.smat_fcwb().to_dataframe().T
ax = sns.heatmap(smat, cbar_kws=dict(label="raw score"))
ax.set_xlabel("distance [um]")
ax.set_ylabel("dot product")
plt.tight_layout()
This scoring matrix works suprisingly well in many cases! However, how appropriate it is for your data depends on a number of factors:
- How big your neurons are (commonly addressed by scaling the distance axis of the built-in score matrix)
- How you have pre-processed your neurons (pruning dendrites, resampling etc.)
- Your actual task (matching left-right pairs, finding lineages etc.)
- How distinct you expect your matches and non-matches to be (e.g. how large a body volume you're drawing neurons from)
Utilities in navis.nbl.smat allow you to train your own score matrix.
Let's first have a look at the relevant classes:
from navis.nbl.smat import Lookup2d, Digitizer, LookupDistDotBuilder
navis.nbl.smat.Lookup2d: this is the lookup table, it can be passed to any nblast-related class or functionnavis.nbl.smat.Digitizer: this converts continuous values into discrete indices which are used to look up score values in an array, we need one for each axis of the lookup tablenavis.nbl.smat.LookupDistDotBuilderis the helper class which generatesnavis.nbl.smat.Lookup2dinstances from training data
First, we need some training data. Let's augment our set of example neurons by randomly mutating them:
import numpy as np
# Use a replicable random number generator
RNG = np.random.default_rng(2021)
def augment_neuron(
nrn: navis.TreeNeuron, scale_sigma=0.1, translation_sigma=50, jitter_sigma=10
):
"""Mutate a neuron by translating, scaling and jittering its nodes."""
nrn = nrn.copy(deepcopy=True)
nrn.name += "_aug"
dims = list("xyz")
coords = nrn.nodes[dims].to_numpy()
# translate whole neuron
coords += RNG.normal(scale=translation_sigma, size=coords.shape[-1])
# jitter individual coordinates
coords += RNG.normal(scale=jitter_sigma, size=coords.shape)
# rescale
mean = np.mean(coords, axis=0)
coords -= mean
coords *= RNG.normal(loc=1.0, scale=scale_sigma)
coords += mean
nrn.nodes[dims] = coords
return nrn
original = list(navis.example_neurons())
jittered = [augment_neuron(n) for n in original]
dotprops = [navis.make_dotprops(n, k=5, resample=False) for n in original + jittered]
matching_pairs = [[idx, idx + len(original)] for idx in range(len(original))]
The score matrix builder needs some neurons as a list of navis.Dotprops objects, and then to know which neurons should match with each other as indices into that list. It's assumed that matches are relatively rare among the total set of possible pairings, so non-matching pairs are drawn randomly (although a non-matching list can be given explicitly).
Then it needs to know where to draw the boundaries between bins in the output lookup table. These can be given explicitly as a list of 2 Digitizers, or can be inferred from the data: bins will be drawn to evenly partition the matching neuron scores.
The resulting Lookup2d can be imported/exported as a pandas.DataFrame for ease of viewing and storing.
builder = LookupDistDotBuilder(
dotprops, matching_pairs, use_alpha=True, seed=2021
).with_bin_counts([8, 5])
smat = builder.build()
as_table = smat.to_dataframe()
as_table
Out:
Drawing non-matching pairs: 0%| | 0/46442 [00:00<?, ?it/s]
Comparing matching pairs: 0%| | 0/10 [00:00<?, ?it/s]
Comparing non-matching pairs: 0%| | 0/11 [00:00<?, ?it/s]
Now that we can have this score matrix, we can use it for a problem which can be solved by NBLAST: we've mixed up a bag of neurons which look very similar to some of our examples, and need to know which they match with.
original_dps = dotprops[: len(original)]
new_dps = [
navis.make_dotprops(augment_neuron(n), k=5, resample=False) for n in original
]
RNG.shuffle(new_dps)
result = navis.nblast(
original_dps,
new_dps,
use_alpha=True,
scores="mean",
normalized=True,
smat=smat,
n_cores=1,
)
result.index = [dp.name for dp in original_dps]
result.columns = [dp.name for dp in new_dps]
result
Out:
Preparing: 0%| | 0/1 [00:00<?, ?it/s]
NBlasting: 0%| | 0/5 [00:00<?, ?it/s]
You can see that while there aren't any particularly good scores (this is a very small amount of training data, and one would normally preprocess the neurons), in each case the original's best match is its augmented partner.
Total running time of the script: ( 0 minutes 0.912 seconds)
Download Python source code: tutorial_nblast_03_smat.py